长链非编码RNA(Long-noncoding RNA,lncRNA)是一类转录本长度大于200bp的非编码RNA,可作为人类基因组中一类重要的调控分子通过多种方式发挥其生物学功能. 近年来的研究表明,lncRNA也可以作为一种竞争性内源性RNA(competing endogenous RNA),与miRNA互作结合,形成ceRNA的竞争机制
以下是我们对数据的整理,通过数据库查询分类、数据库功能及用途、示例结合分析、数据库优化等这四大项,进行阐述和演示数据库的查询和使用,希望对您的实验项目有所帮助
数据库查询分类
① lncRNABase:http://starbase.sysu.edu.cn/mirLncRNA.php
② starbase: http://starbase.sysu.edu.cn/
③ RegRNA2.0:http://regrna2.mbc.nctu.edu.tw/detection.html
数据库功能及用途:
1. lncRNABase
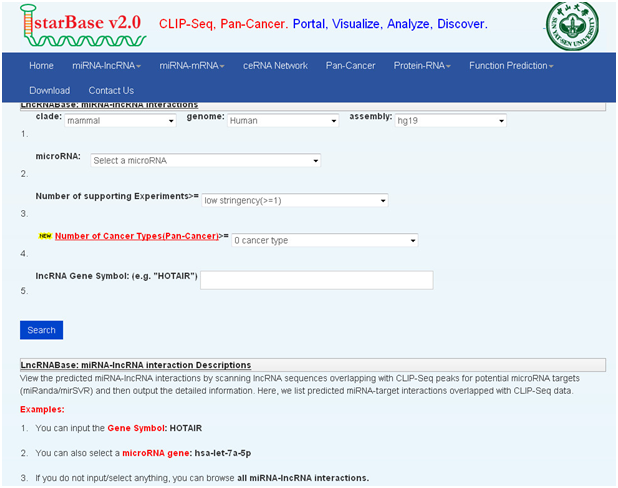
数据库功能:
View the predicted miRNA-lncRNA interactions by scanning lncRNA sequences overlapping with CLIP-Seq peaks for potential microRNA targets (miRanda/mirSVR) and then output the detailed information. Here, we list predicted miRNA-target interactions overlapped with CLIP-Seq data
2. starbase:
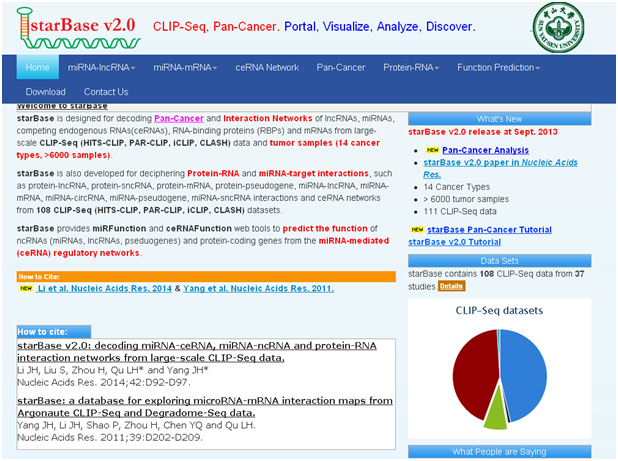
数据库功能:
1. starBase is designed for decoding Pan-Cancer and Interaction Networks of lncRNAs, miRNAs, competing endogenous RNAs(ceRNAs), RNA-binding proteins (RBPs) and mRNAs from large-scale CLIP-Seq (HITS-CLIP, PAR-CLIP, iCLIP, CLASH) data and tumor samples (14 cancer types, >6000 samples).
2. starBase is also developed for deciphering Protein-RNA and miRNA-target interactions, such as protein-lncRNA, protein-sncRNA, protein-mRNA, protein-pseudogene, miRNA-lncRNA, miRNA-mRNA, miRNA-circRNA, miRNA-pseudogene, miRNA-sncRNA interactions and ceRNA networks from 108 CLIP-Seq (HITS-CLIP, PAR-CLIP, iCLIP, CLASH) datasets.
3. starBase provides miRFunction and ceRNAFunction web tools to predict the function of ncRNAs (miRNAs, lncRNAs, pseduogenes) and protein-coding genes from the miRNA-mediated (ceRNA) regulatory networks
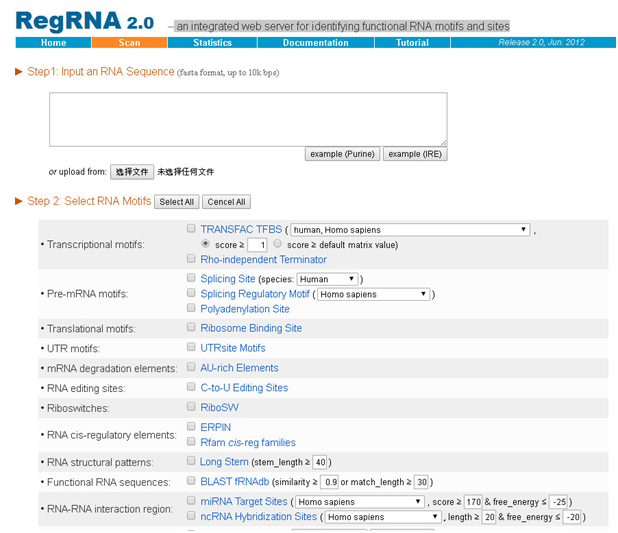
3. RegRNA2.0:
an integrated web server for identifying functional RNA motifs and sites
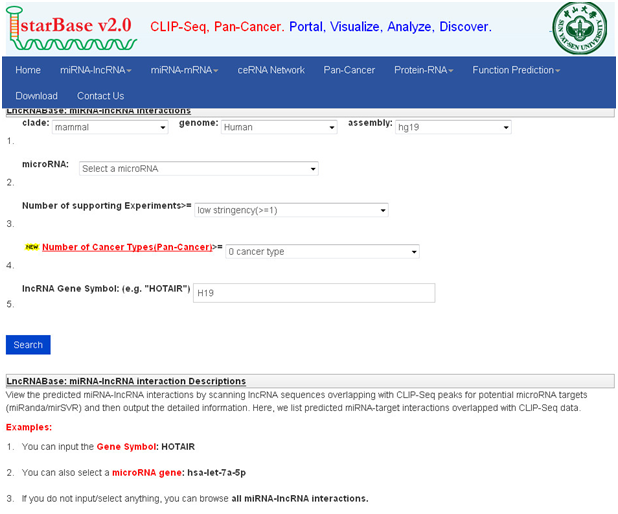
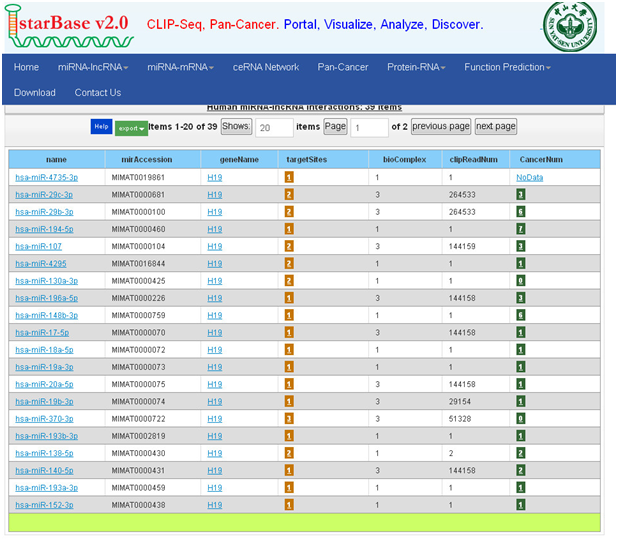
示例结合分析:以H19为例
lncRNABase:
1. 打开主页面
2. 在“lncRNA Gene Symbol”对话框中输入H19
3. 点击“search”,查询即可获取
- 本文固定链接: https://maimengkong.com/kyjc/1695.html
- 转载请注明: : 萌小白 2024年2月21日 于 卖萌控的博客 发表
- 百度已收录
